Reconstructing point forces at focal adhesions
This website provides technical information on a procedure which we developed
in order to calculate localized (point) forces at focal adhesions from elastic substrate
data. Experimental and theoretical issues related to our procedure as well
as results obtained can be found in the following publications:
-
N. Q. Balaban, U. S. Schwarz, D. Riveline, P. Goichberg, G. Tzur, I. Sabanay,
D. Mahalu, S. Safran, A. Bershadsky, L. Addadi and B. Geiger, Force and
focal adhesion assembly: a close relationship studied using elastic micro-patterned
substrates, Nat. Cell Biol. 3: 466-472 (2001)
(abstract,
PDF)
-
U. S. Schwarz, N. Q. Balaban, D. Riveline, A. Bershadsky, B. Geiger, S.
A. Safran, Calculation of forces at focal adhesions from elastic substrate
data: the effect of localized force and the need for regularization, Biophys.
J. 83: 1380-1394 (2002) (abstract,
PDF)
-
U. S. Schwarz, N. Q. Balaban, D. Riveline, L. Addadi, A. Bershadsky, S.
A. Safran, B. Geiger, Measurement of cellular forces at focal adhesions
using elastic micro-patterned substrates, Mat. Sci. Eng. C 23:
387-394 (2003) (abstract,
PDF)
-
B. Sabass, M. L. Gardel, C. Waterman and U. S. Schwarz.
High resolution traction force microscopy based on experimental and computational advances.
Biophys. J., 94:207-220, 2008.
(abstract,
doi:10.1529/biophysj.107.113670,
PDF)
The BPJ 2002 paper gives the technical details as also explained below
and the BPJ 2008 paper compares the results for the reconstruction of
point forces with more general results for a distributed traction
field.
General remarks
Measuring cellular forces on the level of single cell-matrix contacts is
a difficult task for several reasons, and there is no single techniques
yet which avoids all potential problems. Here we list some of the issues
involved:
-
All force sensors will be some kind of spring, whose spring constant has
to be matched to the cellular force. There are several techniques which
have been employed over the last years to measure pN-forces on the level
of single molecules (including atomic force microscopy and optical tweezers),
but all of them are too weak to measure the nN-forces which act at single
cell-matrix contacts. Prestressed thin elastic films turned out to allow
quantitative analysis of the forces exerted by keratocytes, but were too
weak for fibroblast traction, for which one has to use thick elastic films.
-
Typically a cell adheres simultaneously at hundreds or thousands of contacts.
Local force probes like cantilevers micromachined in silicon wafers have
been shown to work for single contacts, but in order to monitor all contacts,
one needs many of those. Microfabricated arrays of elastic needles can
provide sensor densities of down to one per square micron, but restrict contacts
to form on the needle tips.
-
All techniques used up to now are restricted to measure force only in certain
directions: eg a cantilever device or microneedle can only measure force
perpendicular to the lever arm, and elastic substrates have been used only
to measure forces tangentially to the plane of the substrate.
-
Depending on cell type, cells react very sensitively to chemical, mechanical
and topographical features of their environment. Any new experimental technique
introduced has to be proven not to disturb the normal functioning of the
cell. In this vein, the bulky features of cantilevers and microneedles,
the small rigidity of elastic substrates and the topographic patterning
of elastic substrates might turn out to be problematic.
-
The ideal force probe would allow for direct reading of the force value.
For example, a cantilever or microneedle with a known spring constant allows
direct measurement of force by reading of displacement. In contrast, elastic
substrates need calculations to reconstruct the force pattern from the
displacement data.
Up to now, the main technique for measurement of cellular forces has
been the elastic substrate method, which was pioneered by Harris and
coworkers in the early 1980s and turned into a quantitative technique
by Dembo and coworkers in the late 1990s. The elastic substrate method
performs rather favorable in regard to most of the issues listed
above: by tuning the compliance of the elastic medium, one can match
cellular forces; adhesion contacts can be distributed at any number
and at any position over the substrate; force is measured tangentially
to the substrate, but little force is expected to act perpendicular;
elastic substrates are flat and therefore can be easily compared with
results for glass and plastic dishes. The main problem with elastic
substrates is the issue of reading out force: since elastic stress and
strain decays slowly with distance, local information about single
sites of adhesions has to be obtained by non-trivial calculations
extracting a force estimate from the displacement data.
Recently, we adapted the elastic substrate technique for the
special purpose of measuring forces at single focal adhesions. Our
technique combines three elements:
-
The use of micro-patterned elastic substrates allows immediate visualization
of cell traction and easy extraction of displacement. Thus it is an
attractive alternative to the use of marker beads embedded in the substrate.
In our work, we used both topographic and fluorescent patterning, which
can be evaluated by image analysis of phase contrast and fluroescence pictures,
respectively.
-
The use of GFP-vinculin allows to extract position, size and orientation
of the focal adhesions from fluorescence pictures.
-
Numerical solution of the (ill-posed) inverse problem of linear elasticity
allows to estimate the force pattern from the displacement data. In contrast
to earlier work, we assume point forces at focal adhesions, whose positions
are known from the fluorescence data. This procedure has two advantages:
the resulting numerics is much faster than for the case of a continuous
force field underneath the cell body, and the calculated forces can be
easily correlated with other features of the focal adhesions (namely size
and orientation).
Data files
The input and output data of our numerics consist of several ASCII datafiles:
-
lines.dat: displacement data extracted by image processing of phase
contrast or fluorescence pictures showing the patterned substrate. Our
micropattern consisted of a cubic array of small dots, whose positions
had to be determined with and without traction. In order to get the reference
image, cells were trypsinized, but the reference points can also be constructed
by extrapolating the undistorted coordinate system from the far field image
of the image under traction. In order to extract the dot positions, we
used the water algorithm (E. Zamir et al., J. Cell Sci. 1999), which fits
an ellipse to each dot. The ellipse midpoint is then identified with the
dot position. Each line of the file contains data for one dot displacement
in the form x1 y1 x2 y2.
-
focals.dat: focal adhesion data extracted by image processing from
fluorescence pictures. Again the water algorithm has been used to find
focal adhesion positions (moreoever it gives focal adhesion size and orientation,
which were later used for correlation studies). Each line contains data
for the position of one focal adhesion in the form x y.
-
newlines.dat: in principle the same data as in lines.dat, but cleaned
by some data whose use is not reasonable for some reason. For example,
since our approach uses point forces at focal adhesions (ie Green functions
which diverge like 1/r), displacement data should not be used if it is
closer to the focal adhesion midpoint than the typical lateral extension
of focal adhesions. In our paper in BPJ, we showed that with this precaution,
the point forces described by Green functions are good approximations
in the sense of a force multipolar expansion to the distributed forces
acting at focal adhesions.
-
forces.dat: this is the desired output of our calculation: for each
line in newfocals.dat, forces.dat gives the estimate point force in the
form fx fy.
Numerical procedure
As explained in our paper in BPJ, the inverse problem of linear elasticity
(calculating force from displacement) is ill-posed and the calculated force
pattern will be unreliable if not regularized by some appropriate side
constraint. The simplest solution to this problem is use of Tikhonov regularization,
which is a well established technique covered in many textbooks. For our
work, we have used a freely available package of Matlab routines to do
this job. The author of these routines is Prof.
Per Christian Hansen, who has turned his PhD-thesis on this subject
into a book (P. C.
Hansen, Rank-Deficient and Discrete Ill-Posed Problems: Numerical Aspects
of Linear Inversion, SIAM, Philadelphia, 1998). His accompaying software
Regularization
Tools can be downloaded from his webpage and is also available in the
NumerAlgo directory at Netlib. If you want to use our program, you
should first download our routines and
extract them to some directory. Then you should download
Prof. Hansen's software and unpack it into a subdirectory with the
name Hansen. If you then start up Matlab for the directory with
our routines, there should be the data files lines.dat and focals.dat
included in our software bundle to work with. These two files
represent artificial data which we generated in order to check how our
technique is working.
First you should use the routine data.m, which converts the file lines.dat
into newlines.dat. It also produces the following plot:
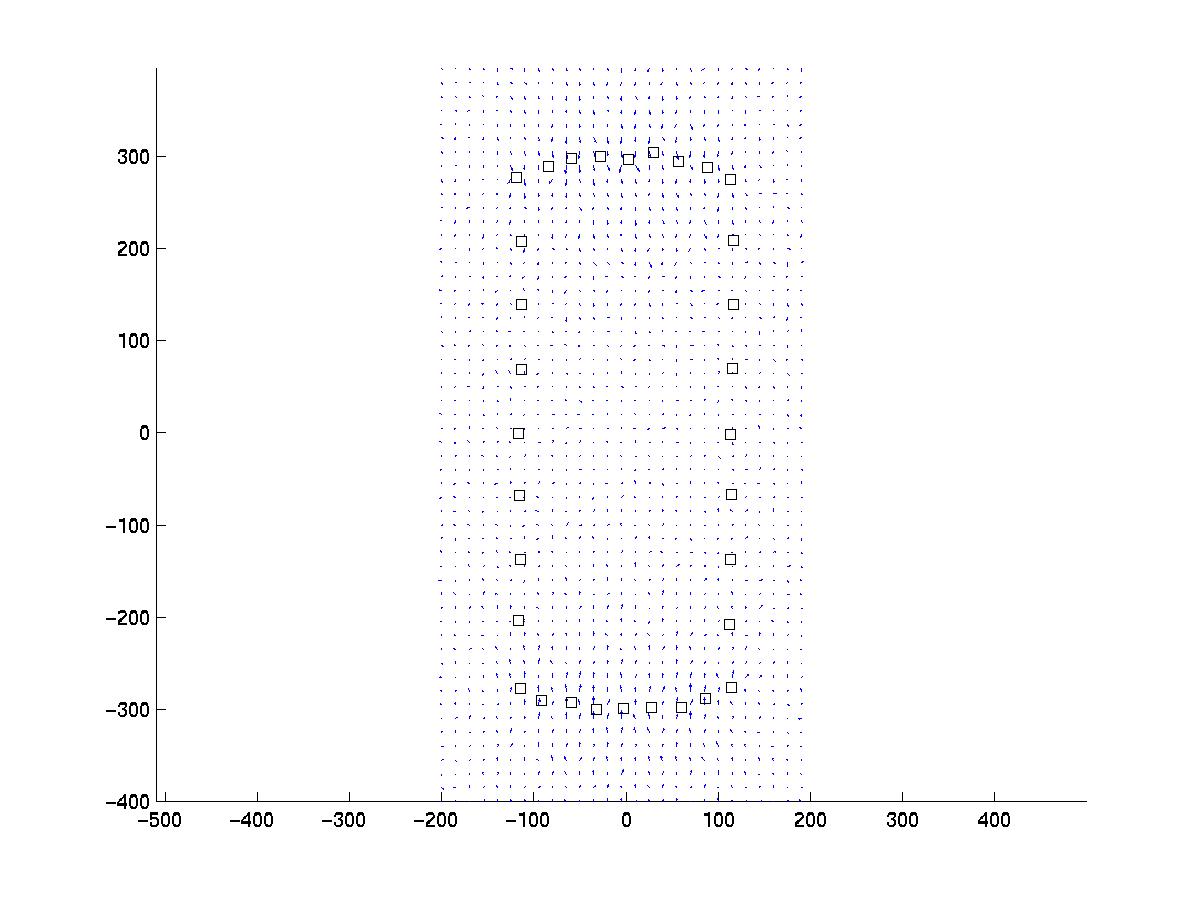
Our artificial pattern of point force positions (squares) can be
imagined to be a set of focal adhesions distributed along the rim of a
polarized cell. The resulting displacement has been calculated on a cubic
array and some noise has been added with a standard deviation of sigma
= 1 pixel, which in an experiment has to be estimated from the data, eg
by analyzing images without cell traction.
Next you can use the routine inverse.m to estimate the original force
pattern from this data. The routine first reads in the data newlines.dat
and focals.dat and plot the same picture as above. It then calculates the
matrix and its singular value decomposition for the linear mapping between
force and displacement from the Green function for an isotropic elastic
halfspace. This might take a while, depending on the number of displacements
and focals. Once this is done, the system of linear equations can be quickly
inverted for different values of the regularization parameter. The program
then plots the residual norm R as a function of regularization parameter
lambda (blue line with sigmoidal shape):
<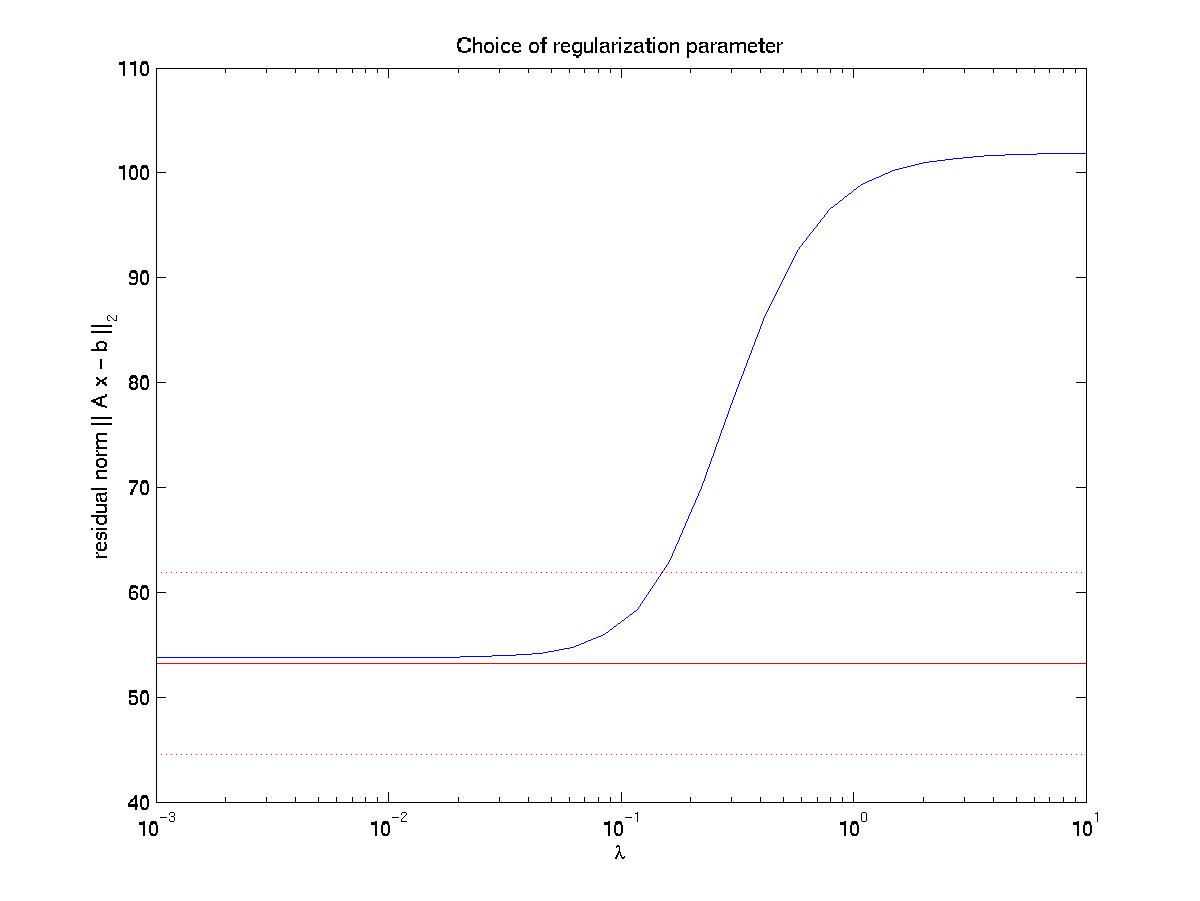
The solid and dotted red lines represent the chi-squared estimate
for the confidence interval for R, based on the value for sigma, the
displacement noise level. For most noise realizations, the solid red
line will cut the blue one, indicating the optimal value for
lambda. In the case shown here, it just fails to do so, thus one has
to choose a value which is closer to the upper end of the
interval. This is a weakness of the chi-square criterion. Another
simple criterion would be to determine the value of lambda at which
the blue curve starts to rise significantly from the plateau, eg as
evidenced by a L-curve plot (not shown here) or by visual
inspection. In our case, this suggests a value around lambda =
0.03. Giving a lambda value to the program yields a plot for the
corresponding force estimate. At the same time, the force estimate is
saved as forces.dat. The program then loops and asks for a different
value of lambda. Giving lambda = 0 terminates the loop. Here we also
show force reconstructions for lambda = 0.01 and 0.1, which bracket
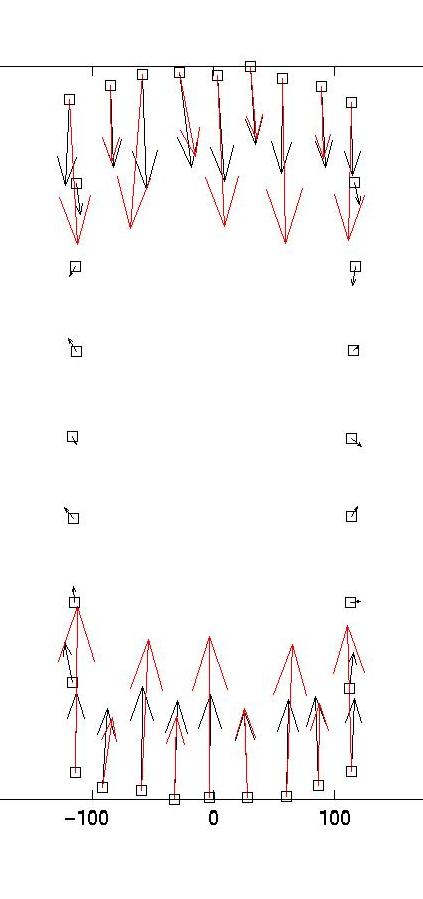
the optimal estimate for lambda = 0.03. The red arrows are the
original forces, which we know since we use an artificial force
pattern:
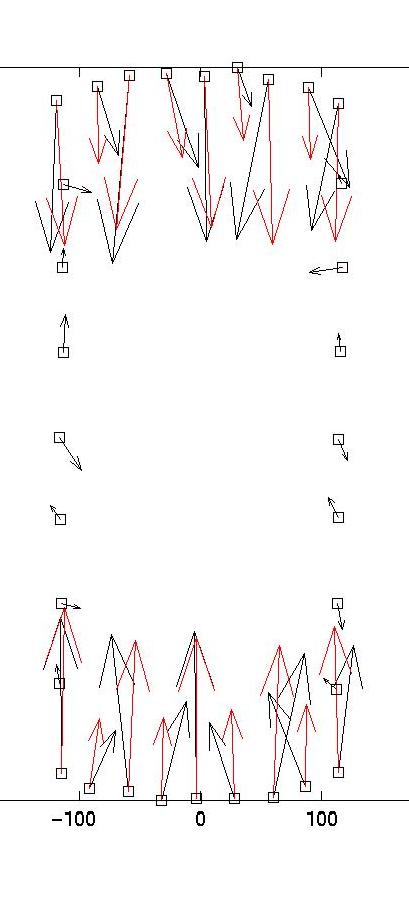
|
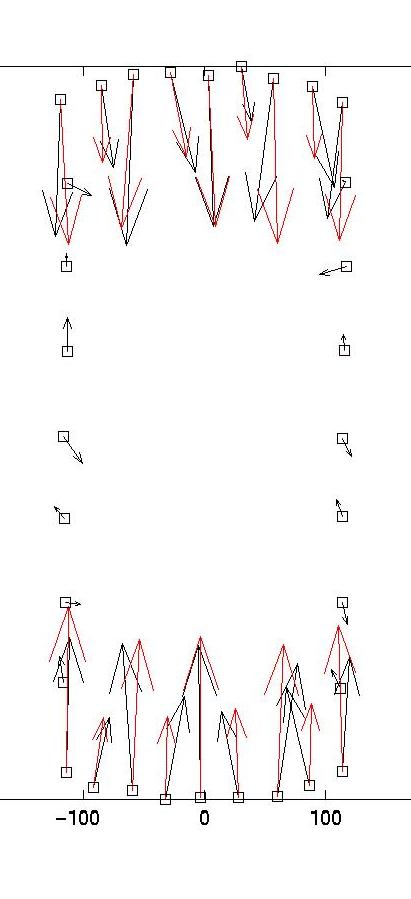
|
|
|
lambda = 0.01 (too small)
|
lambda = 0.03 (optimal)
|
lambda = 0.1 (too large)
|
On the left, we see that lambda values which are too small result
in force details which are artifacts resulting from the noise in the
displacement data (eg original forces had only small x-components and
vanish at the sides). On the right, we see that lambda values which
are too large lead to small forces with little spatial resolution (eg
the magnitude of the original forces oscillates). As explained in our
paper in BPJ, these calculations are estimates and a perfect
reconstruction of the original force pattern cannot be expected. Yet
for the optimal lambda value, the agreement between original and
reconstructed forces is certainly not bad. In particular, the spatial
resolution between the different contacts is reasonable. If one uses
our program on experimental data, one should of course comment out the
few lines related to the original force data, which only exists for
artificial force patterns like the one provided with our software.
Finally one has to calibrate the data. The Green function used in
our software is specified for the incompressible case of Poisson ratio
= 0.5. Moreover one has to know the Young modulus E and the
microscope magnification. Eg for a 100x magnification, 1 pixel =
0.13328 micron. In order to get the force scale right, one has to use
the following formula:
force[nN] = E[kPa] * [pixel/micron]^2
Eg for E = 10 kPa, the factor 10*(0.133)^2 = 0.177 arises: one pixel
correspond to 0.177 nN.
|
This software is free, but if you use it for your own work,
you should acknowledge its use and cite our published work (i.e. the
papers in Nat. Cell Biol. and Biophys. J.). In particular, you should
read the paper in Biophys. J. to understand what the routines are
about. For comments and questions, please address
Ulrich.Schwarz@bioquant.uni-heidelberg.de.
|
Last modified Mon May 4 13:25:57 CEST 2009
by USS.
Back to home page Ulrich Schwarz.





